 Nevertheless, the model of two-state folding does not encompass the breadth of the folding problem under physiological conditions. The combined weight of many folding studies supports the idea that small globular proteins can fold productively in isolation in a two-state fashion. These two observations, namely that small globular proteins attain their native conformations autonomously and that they do so on surprisingly short timescales, serve to frame the biophysical problem of protein folding. As Levinthal famously argued, if the sequence of a peptide did not place some restrictions on the conformational landscape accessible at physiological temperatures, an exhaustive search over all conformational degrees of freedom would take an unreasonably long time. This property, first presented in historic work by Christian Anfinsen, implies that both the native conformation and folding trajectory of a protein are encoded in its primary structure. However, recent work is beginning to shed light on key aspects of chaperonin function, and how their unique properties underlie their contribution to maintaining cellular proteostasis.Ĭhaperonins: The Protein Folding MachinesĪmong the most striking aspects of protein biology is the manner in which polypeptide chains routinely and rapidly attain an active three dimensional structure with high fidelity. A complete structural and mechanistic understanding of this highly conserved and essential chaperonin remains elusive. Folding of many cellular substrates of TRiC cannot be assisted by any other chaperone. TRiC facilitates folding of approximately 10% of the eukaryotic proteome, including many cytoskeletal components and cell cycle regulators. In eukaryotes, the chaperonin TRiC/CCT is hetero-oligomeric, consisting of two stacked rings of eight paralogous subunits each. Group II chaperonins, found in archaea and eukaryotes, contain a built-in lid that opens and closes over the central chamber. Among these, the chaperonins are 1 MDa ring-shaped oligomeric complexes that bind unfolded polypeptides and promote their folding within an isolated chamber in an ATP-dependent manner. 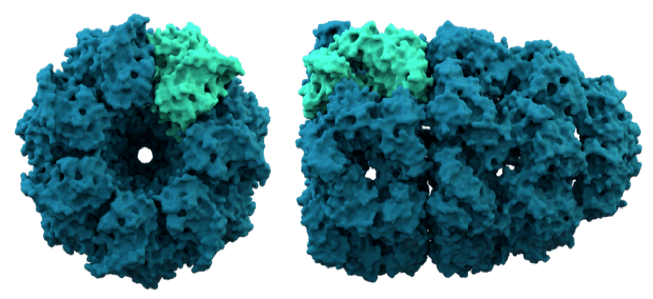
Further refinement of this framework may be necessary because single molecule experiments indicate that there is a great dispersion in the time scales governing the dynamics of the chaperonin cycle.Protein folding in the cell requires the assistance of enzymes collectively called chaperones. Several scenarios for chaperonin-assisted folding emerge depending on the interplay of the time scales governing the cycle. The model shows, in a natural way, that the time scales in the dynamics of the allosteric transitions are intimately coupled to folding rates of the SP. Force-induced unfolding of the SP puts it on a higher free-energy point in the multidimensional energy landscape from which the SP can either reach the native conformation with some probability or be trapped in one of the competing basins of attraction (i.e., the SP undergoes kinetic partitioning). 
We conjecture that the unfolding of SP becomes possible because a stretching force is transmitted to the SP when the GroEL particle undergoes allosteric transitions. A key idea in our framework is that, with coordinated ATP hydrolysis and GroES binding, GroEL participates actively by repeatedly unfolding the substrate protein (SP), provided that it is trapped in one of the misfolded states. However, most of the fundamentals of the GroEL action can be described in terms of the single ring. The architecture of GroEL consists of double toroids stacked back-to-back. We describe a conceptual framework for understanding how the GroEL-GroES system assists misfolded proteins to reach their native states. Molecular chaperones are required to assist folding of a subset of proteins in Escherichia coli. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
